OpenCovid19 Initiative: diagnostic testing through open and collaborative research
Published 11 May 2020 by Elsa Ferreira
Can an inter-communal alliance solve the problem of test shortages? Through the collaborative online laboratory Just One Giant Lab (JOGL), researchers, engineers, designers, start-ups, community managers and public institutions join forces to explore an experimental yet promising diagnostic technique: LAMP.
The global scientific community now has a powerful new tool to mobilise and pool efforts in containing the pandemic: Just One Giant Lab (JOGL), a research and innovation laboratory that is entirely collaborative and open source. Co-founded by Thomas Landrain, founder of La Paillasse open science lab in Paris, along with Marc Santolini and Léo Blondel, JOGL launched in beta in June 2019 in order to tackle the Sustainable Development Goals defined by the United Nations. The Covid-19 crisis further accelerated the expansion of the online platform and community.
On March 1, 2020, the three co-founders and part of the JOGL community launched OpenCovid19 Initiative, a program to “develop open-source and low-cost tools and methodologies” to tackle the crisis. Professional and amateur scientists, open science enthusiasts, students, designers, project managers—a total of over 4,000 individuals are contributing to dedicated projects through the JOGL online platform, Slack channels and Zoom meetings.
In 30 min we will hear the latest developments of the OpenCovid19 initiative!
Come join us on our community call! ?
? https://t.co/Tf9dh8mk0G
? May 6th, 6pm CET pic.twitter.com/3yFVQjPAwY— The OpenCovid19 Initiative ? (@opencovid19) May 6, 2020
To support this process, JOGL is also offering a series of mini-grants of 500-3000 euros to particularly promising projects, thanks to the AXA Research Fund. An “agile” method, explains Thomas Landrain, that allows budding projects to be funded without the risks inherent to significant financial support. If the project is going well, the team can come back to JOGL and apply for more funding.
Among the many ongoing projects are DIY open hardware including syringe pumps, ventilators and bioreactors, educational solutions and quantified health apps. In particular, a number of projects rose to the challenge of making diagnostic tests for Covid-19 cheaper, more efficient and more accessible.
Governments, scientists and the DIY community in the same boat
As progressive lockdown exit strategies are underway, governments are dangerously dealing with serious shortages of testing kits, especially in France, the United States and the United Kingdom. Can open science communities succeed where governments are failing?
“It’s one of these moments in history where an all-hands-on-deck approach is wanted. There can’t be too many people using their brains to find different ways of developing and delivering tests,” advances Ellen Jorgensen, who laid the groundwork for the participative biology movement when she founded Genspace, the first nonprofit community biotech lab in 2009, followed by the public charity Biotech Without Borders.
“We don’t have the resources that governments have,” she continues. “Rolling out a massive amount of tests is not something we can do. A partnership between the DIY community, the traditional scientific community and the government is really the best way to go. That’s exactly what JOGL represents: a way for these different communities to connect, share information and add their strengths to each other.”
LAMP: a proven but neglected technique
The main technique currently being experimented by researchers is LAMP, or Loop-mediated isothermal amplification—a barbaric name for what is actually a much simpler solution than the one currently used to detect Sars-CoV-2, PCR (Polymerase chain reaction). The LAMP technique amplifies the target sequence at a constant temperature in order to detect a pathogen from a sample, whereas PCR relies on thermal cycling, where reactants are exposed to repeated cycles of heating and cooling, a method that requires expensive high-tech laboratory machines.
According to Jorgensen, it’s a proven technique, but one that has not gained wider acceptance because of “barriers to entry” such as user-unfriendly proprietary software, a less “intuitively obvious” reaction than with PCR and, above all, a more sensitive reaction: “The amplification reaction is so efficient that if you open any of the tubes after it happened, that product can contaminate and seed future reactions and give you false positives. If you do contaminate your lab, it might be a month before you can get rid of it.”
LAMP is also less versatile. While PCR can be used quantitatively, LAMP is only used to enhance the presence of a pathogen. “It is a ‘yes’ or ‘no’ result,” says the molecular biologist. “But for Covid-19 it is absolutely perfect. It is a quick and simple diagnostic and can be rolled out quickly and in a variety of different situations.”
#nebLAMPtest : a test in my thermos

One of the essential advantages of LAMP is that the technique doesn’t require any machine, explains Sarah Ware, a biologist involved in the #nebLAMPtest project. She founded two labs in Chicago, including BioBlaze, the first and only community lab in the state of Illinois. She says that the only tools necessary are pipettes and a place that can keep a constant temperature for half an hour, like a water bath or even a thermos—the same kind you use to keep your coffee hot. The cost of materials would be around three dollars.
The #nebLAMPtest team itself encapsulates JOGL’s inter-communal approach: Sarah Ware and BioBlaze from the DIY community; Ellen Jorgensen and her start-up Aanika Biosciences from the private sector; and Chris Monaco, a microbiologist at the U.S. government Centers for Disease Control and Prevention (CDC) health institute.
For this project, the scientists are using results published by New England Biolabs (NEB), a laboratory that produces reagents. A promising report demonstrated that LAMP can be used in conjunction with a PH test, so that results can be confirmed by a simple change in colour, visible to the naked eye, within half an hour. If the reaction remains pink, the test is negative. If it turns yellow, the virus has been detected.
The team is collaborating with partnering labs to reproduce and confirm these findings. The scientists are on the right track but still need to obtain certifications in order to test human patients in their lab. For now, they have been working with an isolated strain of Sars-CoV-2.
#CovidAlert: from GMO to Sars-CoV-2 and beyond

In Oakland, California, Ali Baktas is another molecular biologist closely examining the LAMP technique. His goal is to challenge the centralized process of testing by allowing anybody to do their own detection (a goal widely shared within the biohacking community). “During graduate school and afterwards, my emphasis has been in agriculture,” he says. “I was interested in developing ways for farmers in southern Mexico to monitor their corn against contamination from transgenics.” With a similar ambition for empowering citizens, he has joined the OpenCovid19 Initiative to create an accessible, easy-to-use and shippable molecular test.
According to Baktas, it’s already technically feasible. “There isn’t a lot of novel invention going on, simply we are combining pre-existing, well-established products and techniques,” he says, referring to the LAMP-based backbone of his project. But here too, the team needs to obtain approval, certifications and work on their proof of concept.
Will their solution be ready in time to tackle the current pandemic? “We are working really hard to have it ready as soon as possible. It is not going to be ready next week. A beta version might be ready in a couple of months,” he says, targeting the expected second wave of Covid-19. “Sars-CoV-2 is probably the first of other coming pandemics. There are also other diseases in the world that affect the population for which people need to be able to diagnose themselves cheaply and easily, such as dengue. Sars-CoV-2 is currently the main topic in the world, there is funding and a lot of excitement to work on projects like this, but for us, this is the starting point rather than then end goal.”
#CoronaDetective: the lyophilised test

What about a powder test? The #CoronaDetective project uses a test kit of lyophilised (freeze-dried) reagents, which makes shipping to partner labs a much more streamlined process. Also based on the LAMP technique, this project has partnered with the #nebLAMPtest project and uses GMO Detective, a detector able to ascertain the presence of GMO in food, developed by Guy Aidelberg at the Center for Research and Interdisciplinarity (CRI) in Paris.
Their method is designed to be accessible to anyone. “We can use it in labs, in open labs, and also in institutions such as schools, for instance, to detect the virus on surfaces,” explains Rachel Aronoff, president of Hackuarium, a biohacking association in Switzerland. Practically speaking, she says that the kit will come as eight tubes, with the compounds inside, among which a negative control is included, in order to identify false-positive, which occurs when the testing environment becomes contaminated. To proceed with the test, it will only be necessary to add the liquids, a supplied buffer and the sample, and lay the tubes in hot water. The results will be readable in half an hour through the technique of fluorescence. To make your own DIY fluorescence detector, you can follow instructions supplied for the GMO Detective.
The team works with partners in Chile and Cameroon, among others, ready to test the kit.
#CellFreeSensors: programmable biology
The #CellFreeSensors project uses NASBA (nucleic acid sequence-based amplification), a system of isotherm amplification similar to LAMP that produces amplicons of RNA (ribonucleic acid, chemically close to DNA) to detect a pathogen element.
The team has developed a cell-free system, a technique that can be thought of as programmable liquids, or, according to Vesta Korniakova, a student in applied microbiology and member of the biohacker community in Montreal, as a “coded genetic circuit”.
The case is complex but has the advantage to allow in vitro assay, as opposed as in vivo, a more complicated method that requires working on living organisms that can grow. Korniakova explains: “Anyone who has access to growing bacteria and some materials will be able to follow protocol to make the cell-free machinery required to expressing the generic circuit we are designing”—and thus, when applicable, to visualise the viral genome.
Only requirements: an easy-to-grow E. coli culture and “very easily shareable” DNA primers and sequencers. “As DNA is a very stable molecule, a drop of purified DNA can be sent by post on a paper sheet. More typically, dehydrated DNA is kept in tubes or inside plasmids [DNA molecules], preserved in living bacteria.” Everything else required for the reaction but less accessible, including some enzymes, can be lyophilised and distributed “in bulk”.
The team is spread across three continents, with members in Canada, Chile and Spain.
#DIYBiovsCovid19: Bio hardware
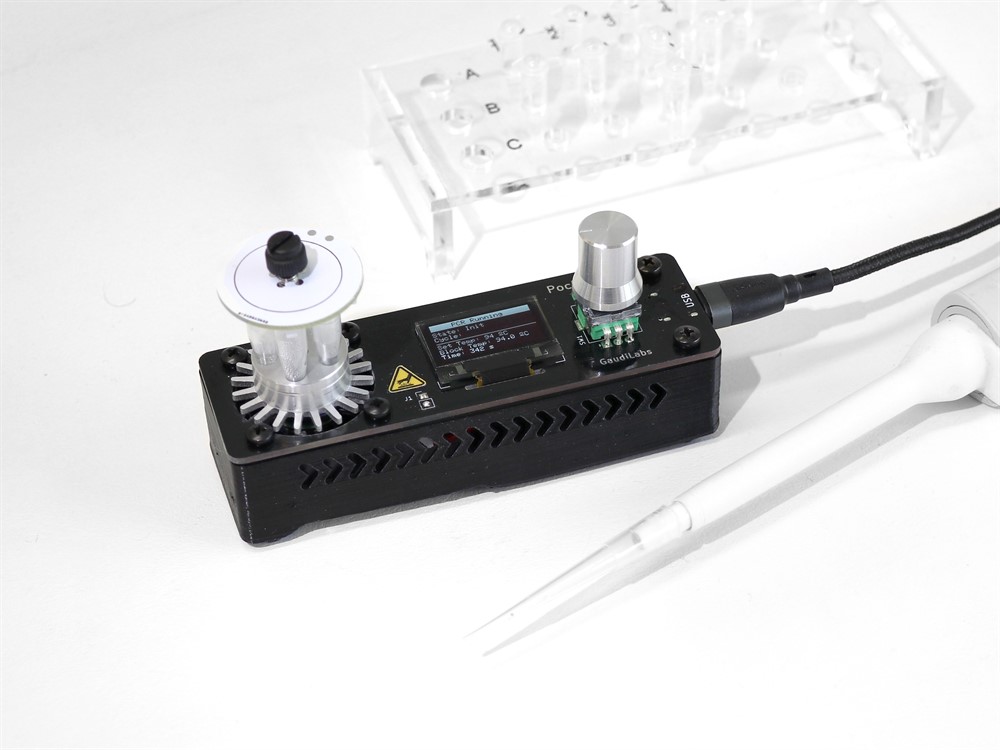

On the hardware shelf is the PocketPCR, developed by Urs Gaudens at his independent research laboratory GaudiLabs. An essential tool for the most common type of Covid-19 diagnostics, PCR amplifies a small sample of DNA or RNA to order to examine it in detail for infectious agents, where the amplification process involves a step-by-step cycle of temperature changes. PCR is also an iconic tool for the open-biology community, says Gaudens, as there have been pioneer projects to reproduce a DIY version of this expensive machine, which can cost several thousand euros.
Gaudens developed several PCR models before refining his pocket version, a heat-block powered by USB. “Since PocketPCR can go through several temperatures, it is easily adapted to the LAMP technique, as it only needs a constant temperature,” he stresses.
Since Gaudens uploaded the project to his website in January, it has drawn interest from many researchers, from Beijing, China to Yogyakarta, Indonesia, but also from companies such as Maternova, which supplies obstetric technologies. Good news, he says, because the higher the volume of orders, the lower the price point. From 99 dollars, he hopes to be able to produce his tool for a third of this price.
At the Starship Factory makerspace in Basel, Switzerland, molecular biologist and self-proclaimed “Swedish mad scientist” Antonio Lamb plans to reproduce the PocketPCR and reduce temperature to the maximum. The resulting hardware can be a complement to the #CoronaDetective project, in conducting the assay under constant temperature, once the powder elements are received, by laboratories worldwide.
“What strikes me from these two first rounds of mini-grants,” concludes Thomas Landrain, “is the incredible quality of the projects.” A quality of proposals that the researcher attributes to JOGL’s signature mode of action: “It fosters the development of stable, sustainable concepts, instead of coming up with quick ideas, like in a hackathon. We allow time for ideas to mingle.” A battle against the pandemic in the form of a marathon, with more rounds to come.
Join the OpenCovid19 Initiative
Submit a project to Round 3 of JOGL micro-grants (by May 18, 6am UTC)
